

International Cancer Genome Consortium (ICGC) – India Project
Introduction
The International Cancer Genome Consortium (ICGC) is one of the most ambitious biomedical research efforts since the Human Genome Project. The ICGC has been launched to coordinate large scale studies to generate high resolution catalogues of genomic alterations in tumors of 50 different cancer types/subtypes that have clinical and societal importance across the globe. These studies undertaken by the member countries of ICGC will eventually analyze over 25,000 cancer genomes at the genomic, epigenomic and transcriptomic levels to reveal the complete repertoire of oncogenic mutations with the goal towards accelerating efforts to develop better ways of diagnosing, treating and preventing cancer. All the ICGC studies share common standards in ethical consent, patient recruitment as well as data collection, storage, analyses and access. Since the overarching goal of ICGC is to maximize public benefit from research, data are being made rapidly available to qualified investigators (www.icgc.org) .

In view of high prevalence and existence of possible interacting environmental factors, India, a founder member of ICGC, has decided to focus on oral squamous cell cancer - gingivo-buccal (OSCC-GB) in the Indian component of the project.
India specific Objectives
-
Create a large collection of appropriate, clinically annotated samples of OSCC-GB.
-
Completely characterize each sample in terms of:
-
All regions of genomic loss or amplification.
-
All mutations in the coding regions of all human genes.
-
All chromosomal rearrangements.
-
Identify all genomic alterations involved in OSCC-GB.
Organization
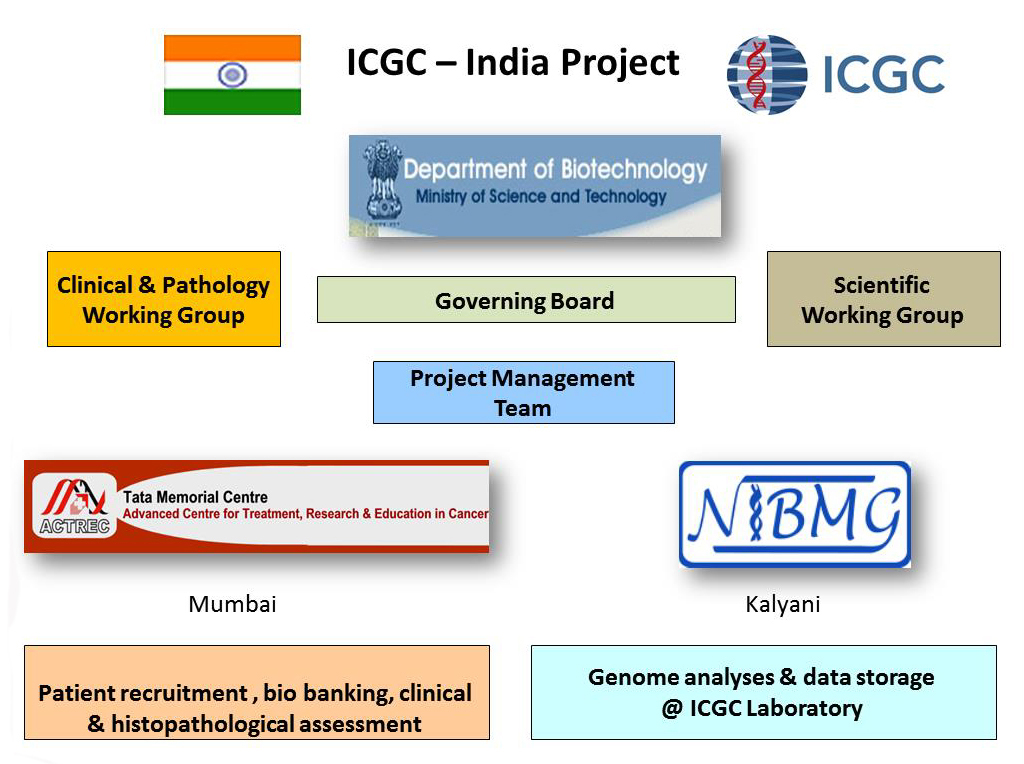
Methods
-
Discovery of single nucleotide and small indel variants (SNVs) by deep exome sequencing of paired tumour and blood DNA from fifty patients of OSCC-GB in Illumina HiSeq-2000 platform.
-
Verification of SNVs discovered in above by deep exome sequencing of paired tumour and blood DNA from these fifty patients in GS-FLX (Roche) platform. Additonal verification by highly multiplexed PCR and deep sequencing in Ion Torrent platform.
-
Analysis of massively parallel sequence data by mapping sequence reads to 1000 Genomes phase 2 reference sequence and variant calling performed by analysis pipeline and Base-by-Base variant caller both developed in NIBMG.
-
Discovery of somatic copy number variations (CNVs) by SNP Chip analysis (2.5 million SNPs) of paired tumour and blood DNA from fifty patients of OSCC-GB in Illumina iScan platform and analysis by ASCAT 2.0.
-
Verification of somatic CNVs by real time PCR in ABI7900 HT platform.
-
Validation of somatic alterations (both SNVs and CNVs) discovered in fifty patients in an independent set of of sixty patients of OSCC-GB by deep exome sequencing and SNP Chip analysis.
-
Presence of HPV and HSV DNA performed in tumour samples by PCR based Sanger sequencing and real time PCR respectively.



